import numpy as np
from spotoptim import SpotOptim
from spotoptim.function import sphere
from spotoptim.plot.visualization import plot_design_points
opt = SpotOptim(
fun=sphere,
bounds=[(-5, 5), (-5, 5)],
n_initial=10,
seed=42,
)
# Generate a Latin-hypercube initial design (in original scale)
X0 = opt.get_initial_design()
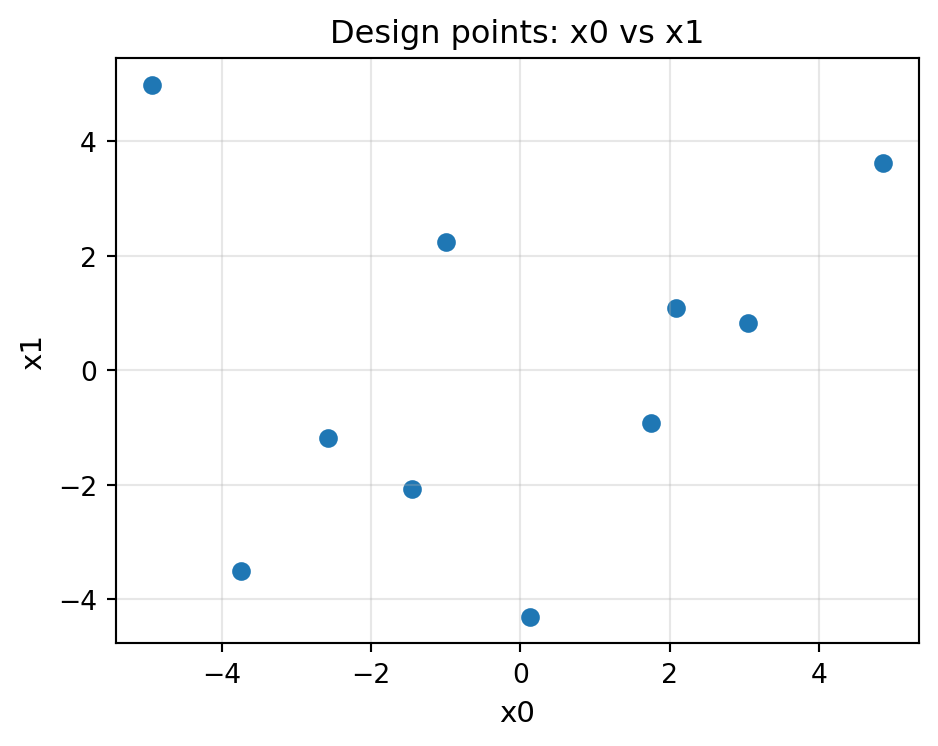
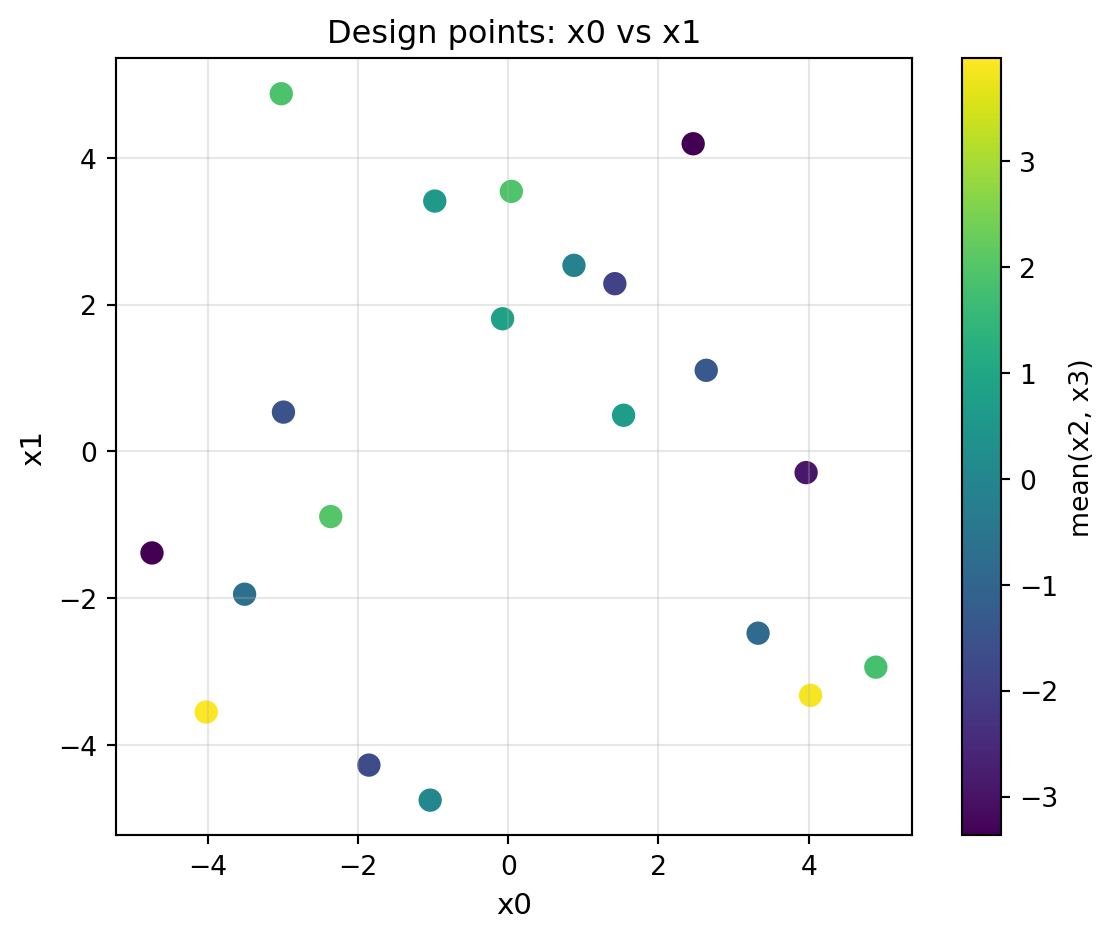
# 2-D design – simple scatter on both dimensions
fig = plot_design_points(X0, i=0, j=1, figsize=(5, 4))